Research
gene editing nucleases
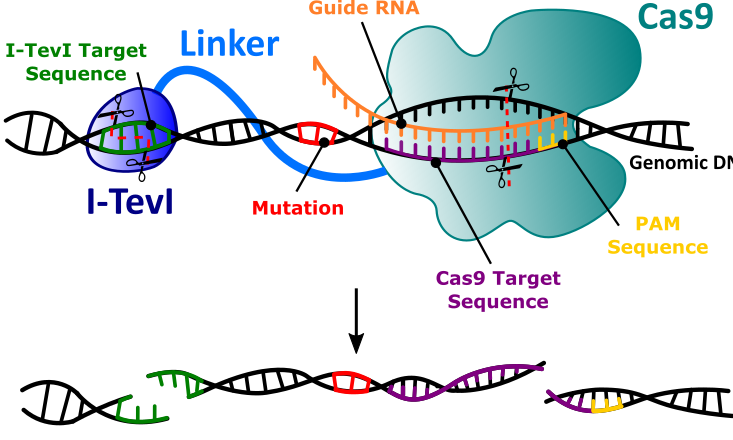
Our lab is actively developing CRISPR-based gene-editing nucleases for targeted modification of genomes. We pioneered the development and use of monomeric nuclease platforms based on the GIY-YIG nuclease domain, I-TevI. Currently, we work on development of dual active-site nucleases that introduce precise, defined-length deletions in genomes. These nucleases are based on fusions of the I-TevI nuclease domain to Cas9 and related proteins. Understanding how I-TevI functions as a site-specific monomeric nuclease to change cleavage specificity is an active line of research in the lab.
microbiome
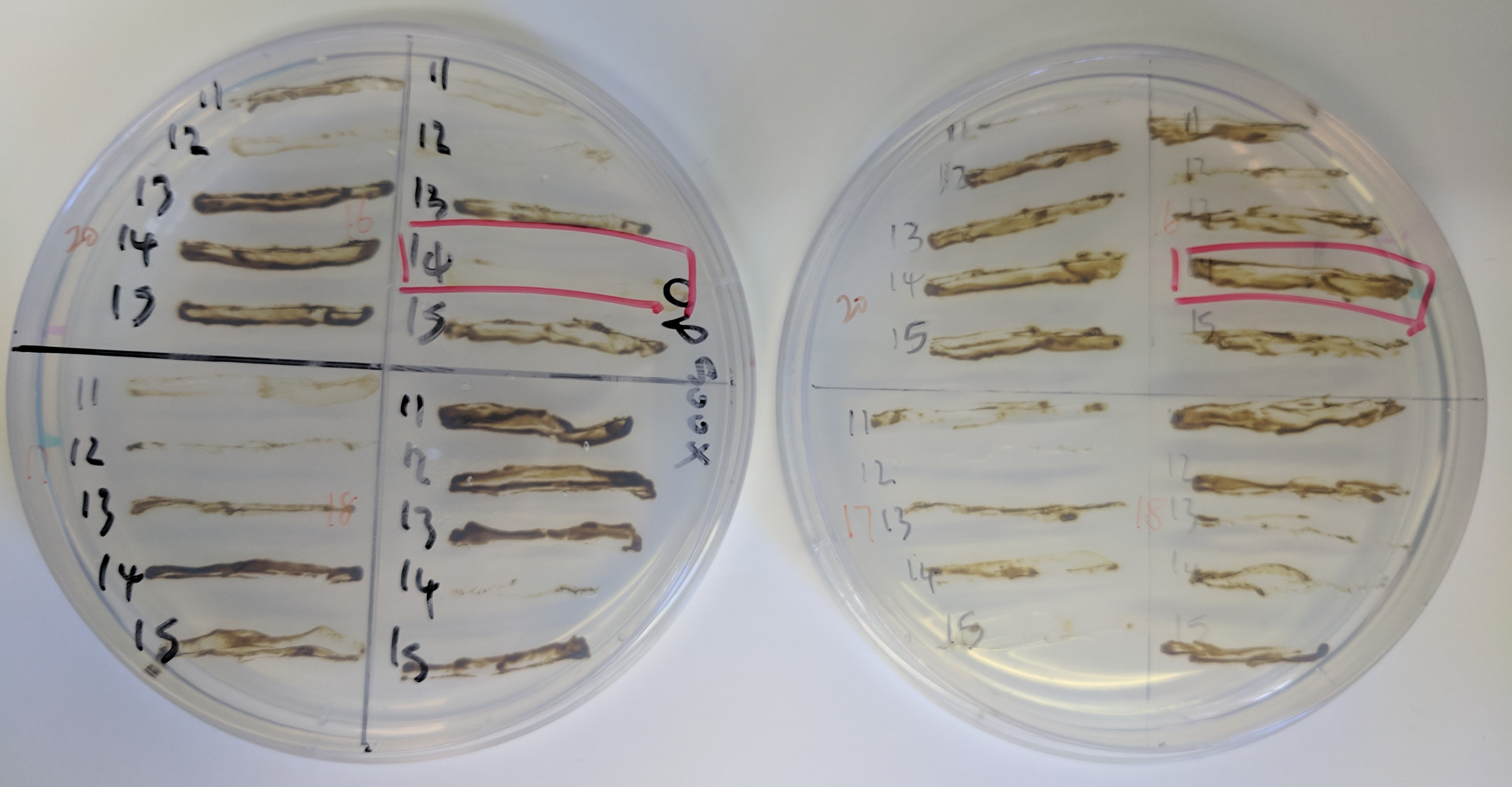
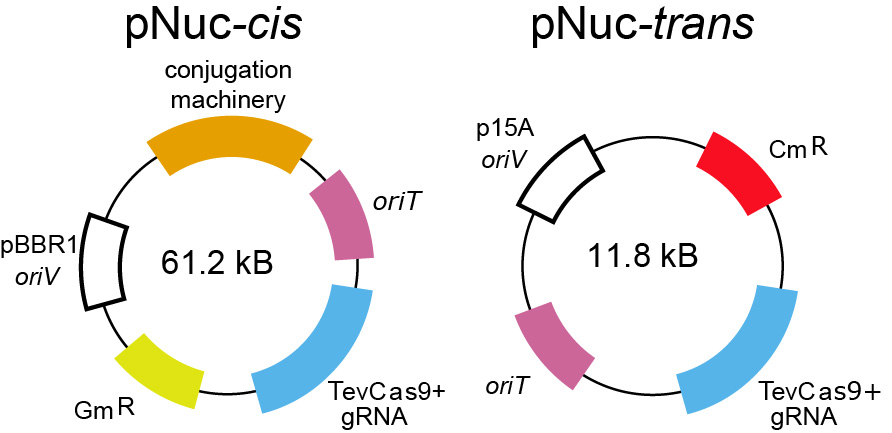
We are developing self-transmissible conjugative plasmids to selectively modulate microbial populations. This area of research is a collaboration with Greg Gloor, Bogumil Karas, David Heinrichs and Emma Allen-Vercoe (Guelph). Our goal is to create broadly transmissible plasmids for delivery of gene products that can modulate microbial populations, either through selective elimination of species by CRISPR-Cas9, or through delivery of gene modules for metabolic manipulation.
synthetic biology
Phaeodactylum tricornutum is a eukaryotic microalgae that is an important model system for synthetic biology and biotechnology. We are developing genetic tools for manipulation of the P. tricornutum genome, including the delivery of Cas9 for gene editing using conjugative plasmids. Our interests lie in using P. tricornutum as a platform for synthesis of value added chemicals and enhanced biofuel production. Currently, we are working on enhancing rates of homologous recombination to facilitate integration of large, synthetic gene clusters.